Gene Detail
Contact
Missing content? – Request curation!
Request curation for specific Genes, Variants, or PubMed publications.
Have questions, comments, or suggestions? - Let us know!
Email us at : ckbsupport@jax.org
| Gene Symbol | STAG2 | ||||||||||
| Synonyms | bA517O1.1 | HPE13 | MKMS | NEDXCF | SA-2 | SA2 | SCC3B | ||||||||||
| Gene Description | STAG2, cohesin subunit SA-2, functions as part of the cohesin complex to regulate spindle formation and sister chromatid association in mitosis (PMID: 12034751, PMID: 24854081, PMID: 29175904). Somatic STAG2 mutations have been reported in glioblastoma, Ewing's sarcoma, bladder cancer, and melanoma, and loss of Stag2 expression has been demonstrated in a variety of other tumor types, including bladder cancer (PMID: 22668012, PMID: 28627627, PMID: 27207471, PMID: 29954776). | ||||||||||
|
|||||||||||
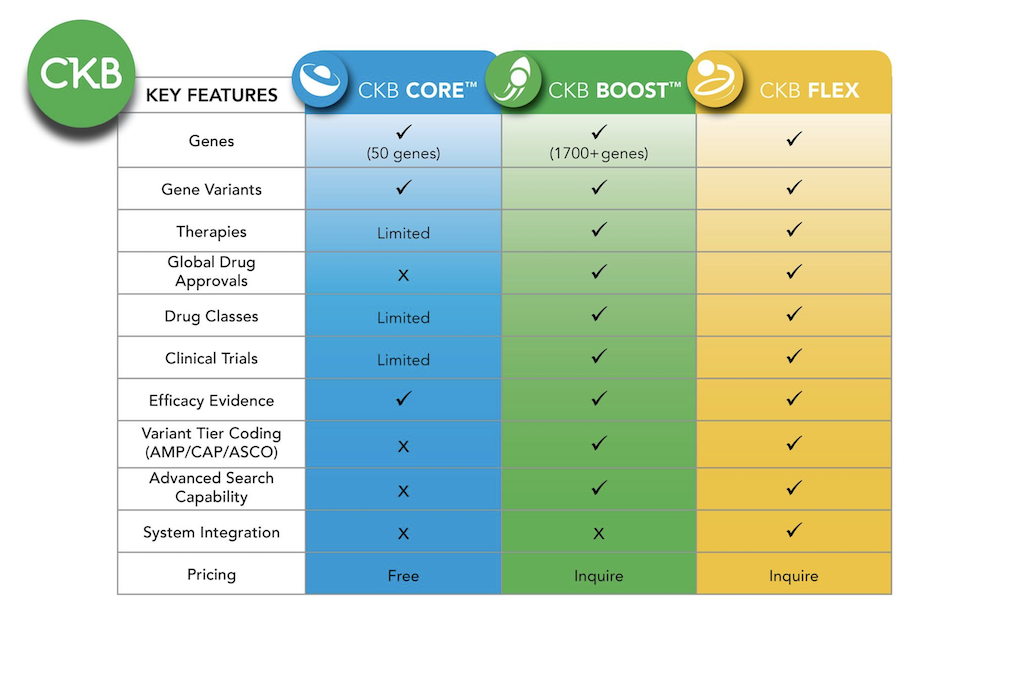
Additional content available in  CKB BOOST
CKB BOOST