Gene Detail
Contact
Missing content? – Request curation!
Request curation for specific Genes, Variants, or PubMed publications.
Have questions, comments, or suggestions? - Let us know!
Email us at : ckbsupport@jax.org
| Gene Symbol | ARID1A | ||||||||||
| Synonyms | B120 | BAF250 | BAF250a | BM029 | C1orf4 | CSS2 | ELD | hELD | hOSA1 | MRD14 | OSA1 | P270 | SMARCF1 | ||||||||||
| Gene Description | ARID1A, AT-rich interaction domain 1A, is a member of the cBaF subunit (PMID: 32303701) of the SWI/SNF chromatin remodeling complex and is involved in cell-cycle activation (PMID: 29136504). ARID1A has been reported to influence PI3K/AKT pathways (PMID: 24618703), and loss of function is commonly found in ovarian clear cell carcinoma (PMID: 32020380, PMID: 32027624), gastric, colorectal (PMID: 28937020), and bladder cancers (PMID: 28583311), while in liver cancer, Arida1a has a context dependent role (PMID: 29136504) and ARID1A promoter hypermethylation has been observed in squamous cell carcinoma (PMID: 32015157). | ||||||||||
|
|||||||||||
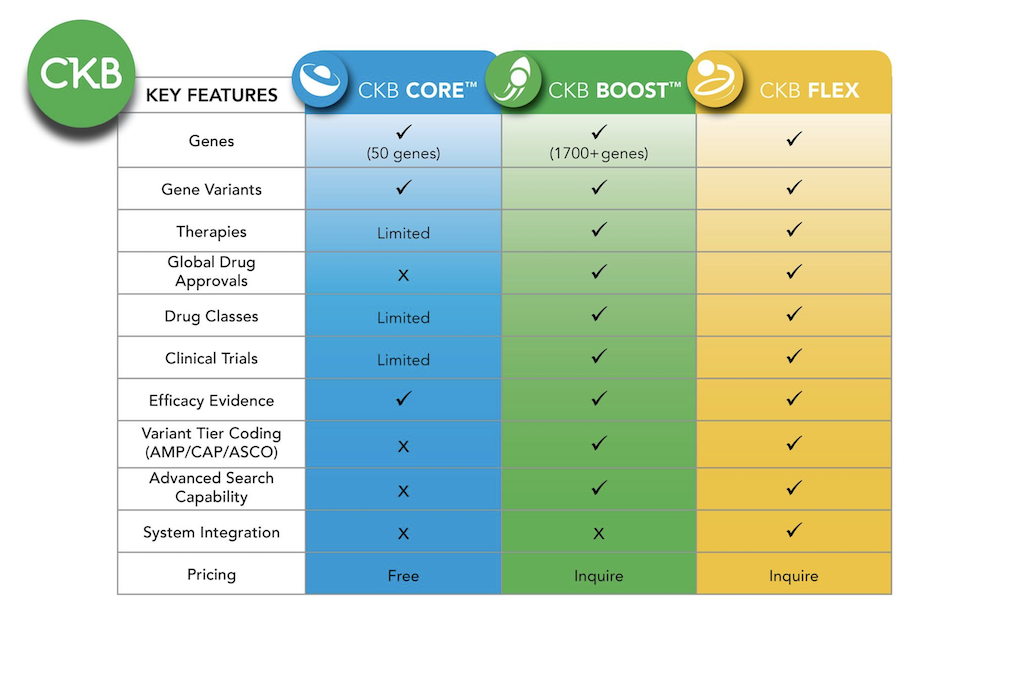
Additional content available in  CKB BOOST
CKB BOOST